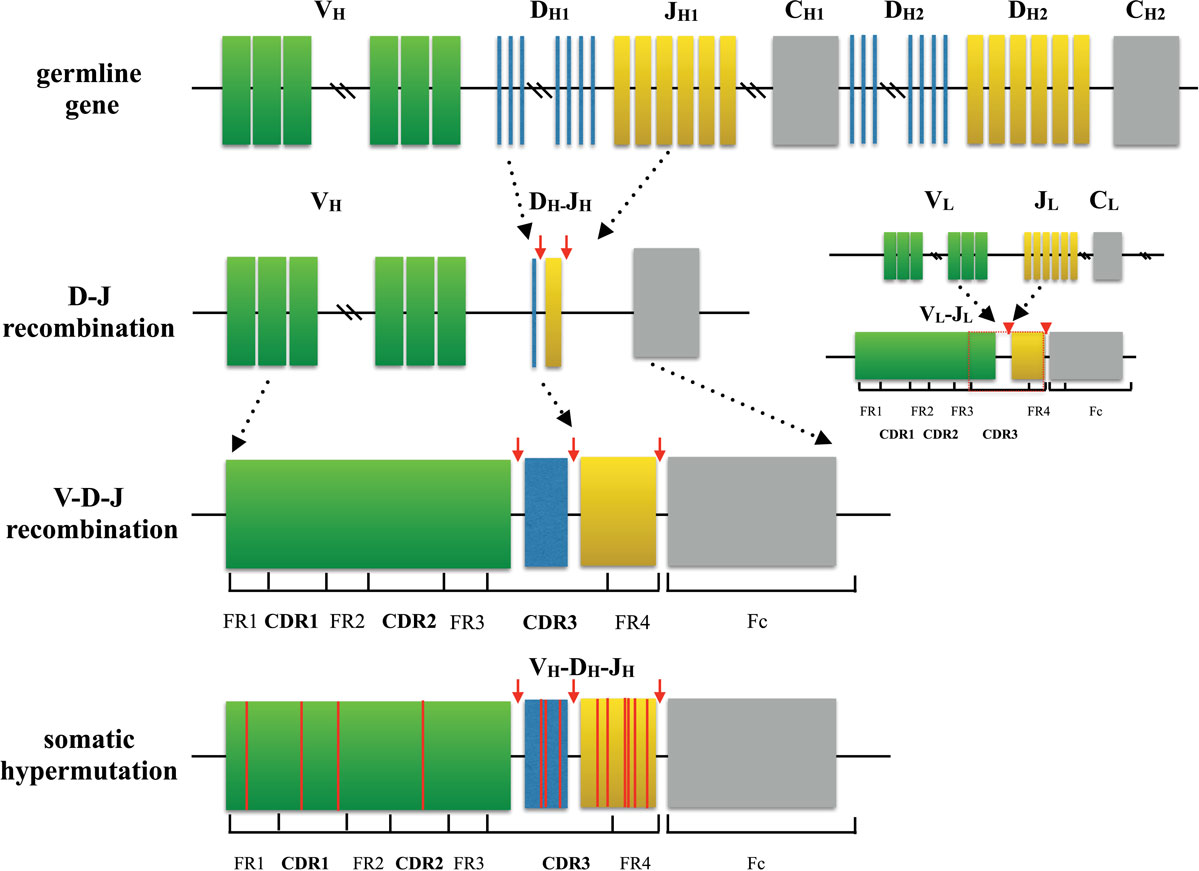
I did a talk Unraveling the Mystery of Immunity at Tedx recently to try to articulate some ideas around Human Immunome, although it was sort of pitched in the context of infectious diseases and vaccines, and I described but did not explicitly name the Human Immunome Program work.Atreca and Bill & Melinda Gates Foundation have collaborated to accelerate the discovery and development of new vaccines and therapeutics for human infectious diseases using Immune Repertoire Capture technology. It is apparent we will need to deliver some early wins in comparing disease state immunizes with healthy for the public to see the potential. But most people want to know the more immediate impact for them (“will this inform cancer immunotherapy” “how will this impact MS, or IBD, or allergy, etc etc”. To me it is obvious that Human Genome > Precision Medicine and other initiatives, and Human Immunome will > same magnitude of impact. In general, I have found that members of the public (and the scientific community for that matter) don’t have a vision for how the complete Human Immunome might impact their lives or work. We have spent the last year interacting with major donors to finance the longer term aspects of the project. The goal is to identify all of the expressed BCR and TCR on the planet. We are slated to sequence complete BCR and TCR repertoires for 1,000 people, and have already started the process. Yes, our laboratory is leading the Human Immunome Program in the Human Vaccines Project network. While currently the number of sequences of known specificity is fairly limited, I think this will rapidly grow as more people apply RepSeq to various disease/vaccination cohorts, and we still may be able to get a rough idea.ĭoes anyone know of any immune repertoire startups already doing this kind of thing? If we could get people interested in having their immune repertoires sequenced, it would then be a great research resource too. 
Relies on having a database of previously described antigen-specific sequences with which to annotate the datasets (see this thread). 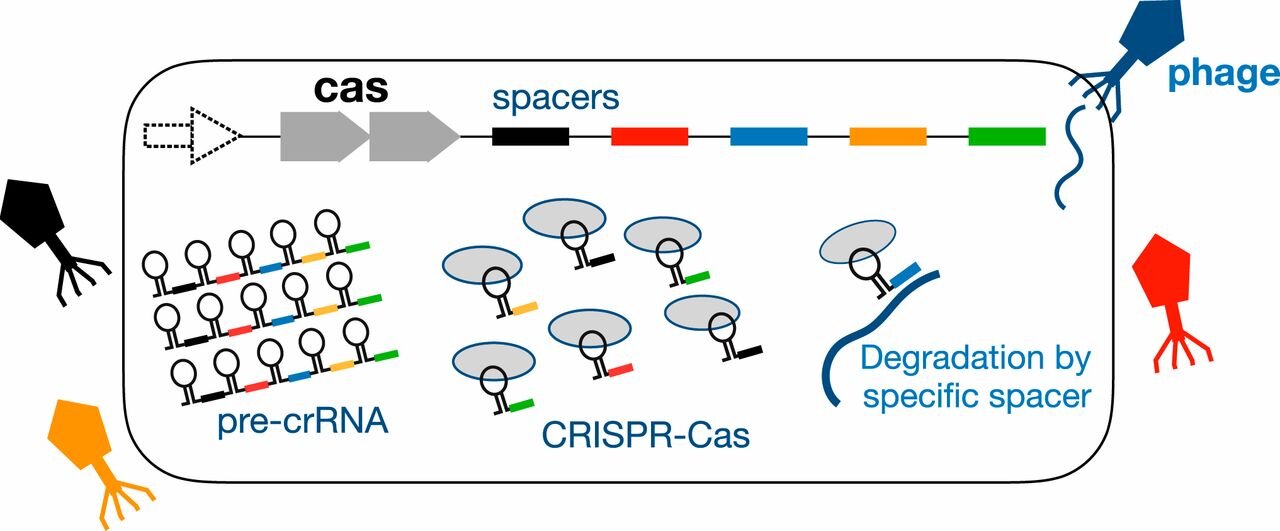
What pathogens have you been exposed to? This is probably the most interesting, but also the most difficult at the moment.How mature is your repertoire? We know how some repertoire features change with age, and would be fun to see if we could predict the age of the people based on their repertoire features alone.Need a good baseline dataset of healthy people so we can then see how people compare to this. How does your repertoire compare to others? PCA of broad repertoire characteristics and where you fit (diversity, CDR3 length, mutation, VJ, etc).Even if it doesn’t say much personal, it is something that looks fun, and illustrates how diverse the repertoire is. Your repertoire is diverse! I find that people’s eyes are always drawn to a nice network representation of a repertoire.A few ideas below, and what would be useful to do to help achieve them. 
I think eye-catching visualisations are the key, even if we don’t fully know the implications of different repertoire features at this point. We could generate similar reports to the one shown above. The problem will be that the samples are relatively sparse (due to limits of blood draws), so good inference is vital. Obviously, some of this can be done with serology, but B cells make it vivid and personal.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
